Xenium iST
Imaging-based spatial transcriptomics (iST) using the 10X Genomics Xenium. Profiles RNA and multiplexed protein in cells and tissues with subcellular resolution.
Do not order Xenium reagents. ATG-C purchases all Xenium reagents through a blanket purchase order. We do not run externally purchased reagents.
Workflow
-
Request a project
We're happy to discuss your project with you. Start your Xenium iST project by requesting a service through iLab or emailing genome@columbia.edu (mention Xenium project in your subject line).
-
Complete the project information form
Fill out this Google Form. You will need your iLab Service ID (format example: GECF(SB)-AB-12), tissue type, panel selection, and number of slides.
-
Pick up Xenium slides
Schedule a time by emailing genome@columbia.edu. Bring a container with dry ice for transport.
-
Prepare your slides
Carefully follow the 10X Genomics Tissue Preparation Guide for FFPE or fresh frozen samples. It is essential that your samples are properly fixed (or frozen), sectioned, and placed on the slide.
-
Return slides to ATG-C
Observe the drop-off times: FF by 4:00 PM, FFPE by 12:00 PM (ATG-C incubates) or 4:00 PM (user incubates).
-
Download your data
We will contact you when the data is ready to transfer. Data must be downloaded within 30 days. Analyze with Xenium Explorer, Xenium Ranger, or Squidpy. Columbia analysis support available on request.
Protocols & Resources
- 10X Genomics Xenium Platform
- FFPE Tissue Preparation Protocol
- Fresh Frozen Tissue Preparation Protocol
- Custom Gene Panel Ordering
- Analysis Software Overview
- Xenium Explorer
- Xenium Ranger
- Squidpy
- Continuing Your Journey After Xenium Analyzer
Analysis Support
The Columbia Genome Center's Single-Cell Analysis Team can help with downstream analysis of Xenium data. Contact genome@columbia.edu and ask about analysis help on a Xenium project.
Frequently Asked Questions
What causes run failures or low-quality data?
The most common cause is low-quality tissue sections. Closely follow the 10X Genomics tissue preparation protocols for FFPE and fresh frozen samples. Fixation problems and poor sample quality are the primary risk factors. See also the 10X KB articles on FFPE fixation recommendations, mRNA degradation prevention, and tissue block compatibility.
How should I assess sample quality before submission?
We recommend H&E staining, DAPI staining, and DV200 assessment for all samples. 10X Genomics recommends DV200 ≥30% as a minimum threshold, though there is not a strong correlation between RIN or DV200 and run quality. See the 10X KB articles on tissue block compatibility, RNA quality assessment importance, and the Tissue & Sample Preparation section.
How many genes are in the pre-made panels?
Human tissue panels range from 266–377 genes. Mouse tissue panels range from 247–379 genes. See the Xenium panels product page for the full catalog. Cell-type annotations are available on the manufacturer's website.
Can I design a custom gene panel?
Yes. 10X Genomics will work with you to choose the genes you want through their panel designer portal. See also the panel designer inputs page. Most designs are automated. Some require advanced design for additional fees.
Can leftover custom panel reagent be stored?
Yes. Leftover custom panel can be stored frozen and should be used within one year following resuspension. See the 10X KB article on probe shipping and storage temperatures.
Are there genes that cannot be included in panels?
Highly expressed RNAs, mitochondrial genes, ribosomal genes, and HLA class 1 genes are discouraged due to optical crowding constraints. Genes with high sequence homology may also be difficult to include. See the panel design getting-started guide for details.
What section thickness is required?
10 μm for fresh frozen and 5 μm for FFPE. For guidance on working with larger tissue sections, see the 10X KB articles on large tissue sections and fiducials coverage.
Should I use fresh frozen or FFPE?
Good results can be obtained with either. Fresh frozen samples tend to have more transcripts per cell; FFPE has better morphology and segmentation.
Who places tissue sections on the slides?
Users place sections themselves or can work with the CUIMC molecular pathology core for sectioning services. Refer to the 10X Genomics tissue preparation protocols for FFPE and fresh frozen samples.
Does ATG-C handle fixation and deparaffinization?
Yes. ATG-C will fix your frozen samples or deparaffinize your FFPE sample slides.
How many tissue sections can fit on one slide?
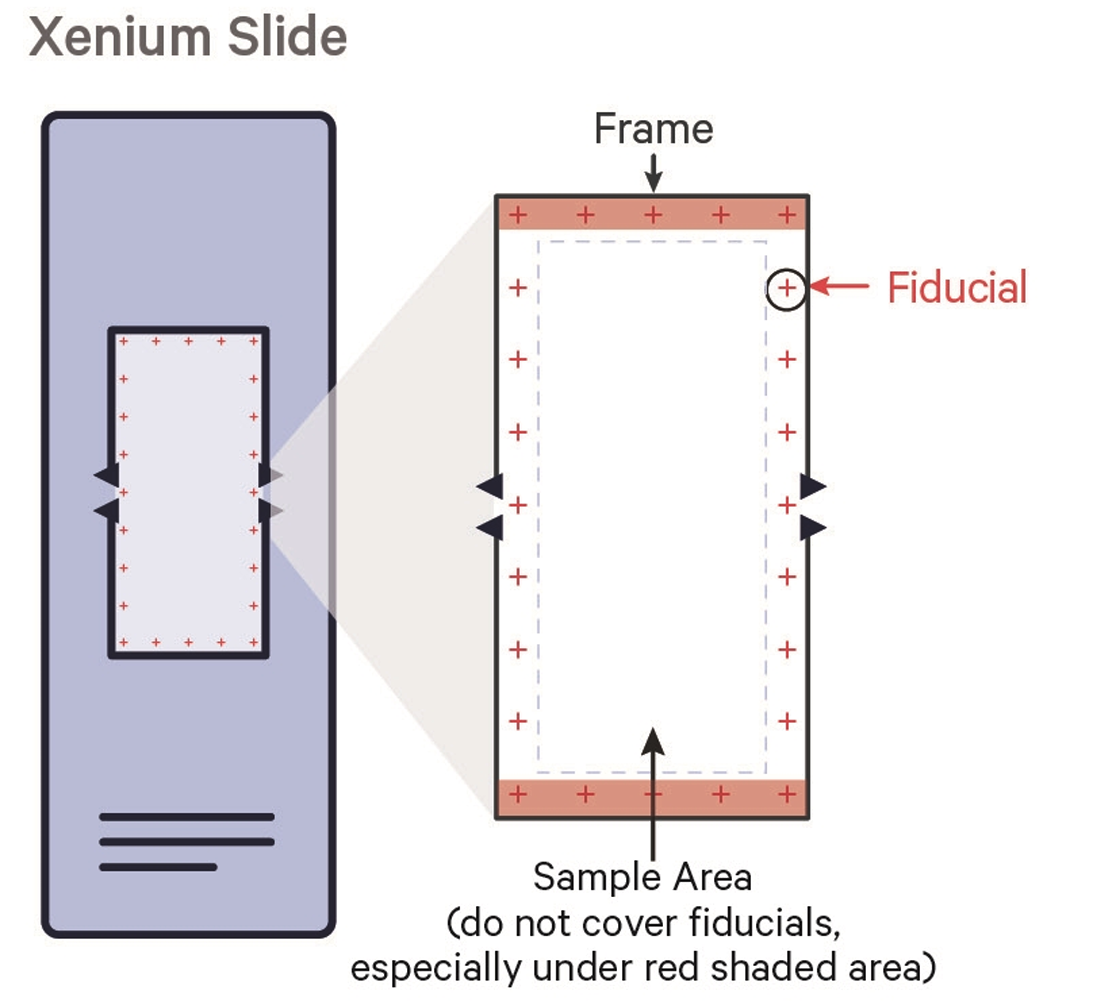
The imaging area is 10 mm × 22 mm per slide. The Xenium recognizes up to 8 regions of interest (ROIs) per slide. Fiducial crosshair markers at the corners must not be covered by tissue. More than 8 samples can be included on one slide with manual demultiplexing post-run (notify ATG-C beforehand). See the 10X KB article on fiducials coverage.
How should FFPE slides be transported after sectioning?
Transport FFPE slides at room temperature immediately after the drying step in a sealed slide mailer or resealable bag.
What should I do with FFPE slides after placing tissue?
Follow protocol: dry upright at room temperature (~10 min with fan, 30 min without), then incubate 3 hours at 42 °C. ATG-C can perform the incubation if slides are delivered by 12:00 PM. See the 10X KB article on FFPE slide storage.
How should fresh frozen slides be stored before submission?
Refer to the 10X KB article on fresh frozen slide storage for storage recommendations.
What are the drop-off times?
Fresh frozen samples: by 4:00 PM. FFPE (ATG-C incubates): by 12:00 PM. FFPE (user incubates at 42 °C for 3 hours): by 4:00 PM.
Can I get my slides back after the run?
Yes. The Xenium Analyzer uses non-destructive analysis and gentle chemistry. Slides can be stained for H&E or immunofluorescence afterward. Notify ATG-C that you want your slides returned and pick them up within 1 business day of run completion.
How large is the output data?
Output data is approximately 50–100 GB per run. ATG-C will provide a download link. Data must be downloaded within 30 days.
What format is the output data?
What are the optics specifications?
The Xenium Analyzer uses a high-NA optics, fast area-scan camera system. Pixel size: 0.2125 μm/pixel. Transcript XY-localization precision <30 nm. Z-localization precision <100 nm. See the 10X KB article on Xenium Analyzer optics and the instrument page.
Is there a specific way to align tissue sections on the slide?
As long as sections are within the fiducial marks, there is no required or recommended alignment. Place tissue wherever it fits within the imaging area. Uniform alignment may facilitate sample comparison.